We are a research group at the Guangzhou National Laboratory. We are a research group at the Guangzhou Laboratory. If you are interested in working with us, please see more information on (Vacancies).
RNA centre (Miao lab) is a computational biology laboratory, which focuses on the research of RNA structure, function ( RNA structural informatics ) and single-cell omics sequencing. We develop new computaitonal approaches (algorithms, databases, integrated computational workflows) to understand the RNA function at regulation level and structure level.
RNA structural informatics:
Our lab seeks an agile and predictive understanding of how RNAs code for information processing and replication in living systems. We are creating new computational and chemical tools to enable the precise modeling and design of these RNAs.
single-cell omics:
We have a longstanding interest in understanding global principles of gene regulation and protein-RNA interactions. We use state-of-the-art genomics approaches, including multi-modal single cell genomics and spatial genomics in combination with machine learning methods to advance our knowledge of cells and tissues.
We are looking for passionate Associate Investigators, Postdocs, Assistant Investigators and Research Assistants to join the team (more info) !
We are grateful for funding from Guangzhou Laboratory (Gzlab), MOST and NSFC .

31. Mar 2026
We published a paper in GigaScience scDenorm: a denormalisation tool for integrating single-cell transcriptomics data. We developed scDenorm, a method to recover raw counts from normalized single-cell RNA-seq data and reduce inconsistencies across datasets. By reversing normalization and rescaling expression values, scDenorm improves data integration and downstream analyses, providing an efficient alternative to reprocessing raw sequencing data.
4. Feb 2026
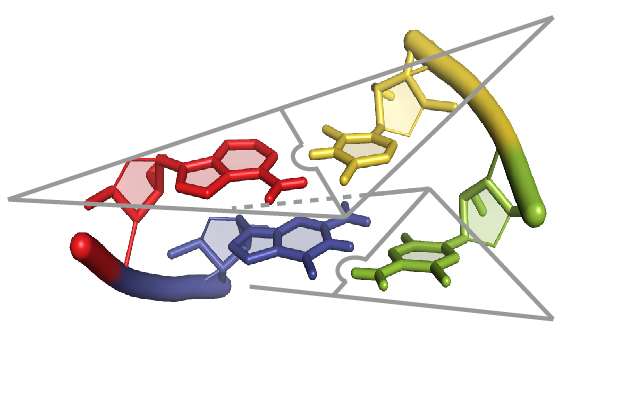
We published a paper in Journal of Biological Chemistry Structural Basis of DNA Aptamer A58 Targeting the N-terminal Domain of Sarbecoviruses Nucleocapsid Protein. We determined the crystal structure of DNA aptamer A58 in complex with the N-terminal domain of SARS-CoV-2 nucleocapsid protein, revealing a unique three-tiered stem-loop binding mode. A58 inhibits N-NTD’s interaction with viral RNA and shows broad-spectrum activity against N proteins from various sarbecoviruses, highlighting its potential as an antiviral agent.
17. Dec 2025
RNAcentral Consortium published a paper in Nucleic acids research RNAcentral in 2026: genes and literature integration. RNAcentral has developed two important new features. First, it integrates literature, introducing LitScan and LitSumm, effectively connecting sequence data with functional knowledge scattered across thousands of articles. Second, RNAcentral introduces gene-level entries, integrating related transcripts into a gene-centric view. Our ribozyme, riboswitches, and RNA aptamers databases directly contribute to the RNAcentral comprehensive database, systematically summarizing the sequence-structure-function relationships of these three RNA types, providing data corpus for AI model training.
1. Dec 2025
We published a paper in Briefings in bioinformatics Systematic identification and characterization of virus lncRNAs suggests extensive structural mimicry of host lncRNAs. We systematically identified vlncRNAs for more than 20 viral species by mining viral infection-related TGS data and further characterized their structure and function. A database named vlncRNAbase was further built to store and organize these newly identified and known vlncRNAs. It would greatly facilitate further research on vlncRNAs in the field.
22. Nov 2025
We published a paper in Nucleic acids research GlycoRNAdb: a database of glycoRNA sequences, structures, abundance, and glycan information across tissues and cell lines. We developed GlycoRNAdb database, the first comprehensive database integrating GlycoRNA sequences, structures, glycosylation sites, expression profiles, and glycan annotations across human/mouse tissues and cell lines, providing an integrative overview of this information.
14. Nov 2025
We published a paper in International journal of infectious diseases Temporal single-cell landscape suggests infection-induced IgM plasma cells as a correlate of durable influenza immunity. We identify an infection-induced IgM⁺ plasma cell population with sustained HLA-II expression, absent after vaccination. These cells may contribute to long-lived immunity and represent a potential target for improving vaccine durability.
21. Oct 2025
We published a paper in Nucleic acids research Ribocentre-aptamer: an integrative, structure-focused database for RNA aptamers. We have launched the Ribocentre-aptamer database to capture the molecular mechanisms of RNA aptamer binding. Our resource systematically curates 191 aptamer types from 669 publications, integrates 510 sequences, 123 small molecule ligands, and 344 protein ligands, and provides experimentally resolved secondary and tertiary structures, ligand binding pockets, affinity metrics, and functional annotations.
15. Oct 2025
Mengchang Xu is joining the lab as a Research Assistant.
3. Oct 2025
We published a paper in Proteins The RNA-Puzzles Assessments of RNA-Only Targets in CASP16. We applied the automated evaluation system accumulated by the RNA-Puzzles community since 2011 to the RNA competition in CASP16 (2024). We systematically compared the consistency between predicted and measured three-dimensional structures, looking not only at global indicators (RMSD, TM-score, GDT, lDDT) but also at key details such as stereochemistry, Watson-Crick/non-Watson-Crick pairing, and base stacking. This revealed that large-size/multimeric RNA and atypical base pairing remain weak points in current methods.